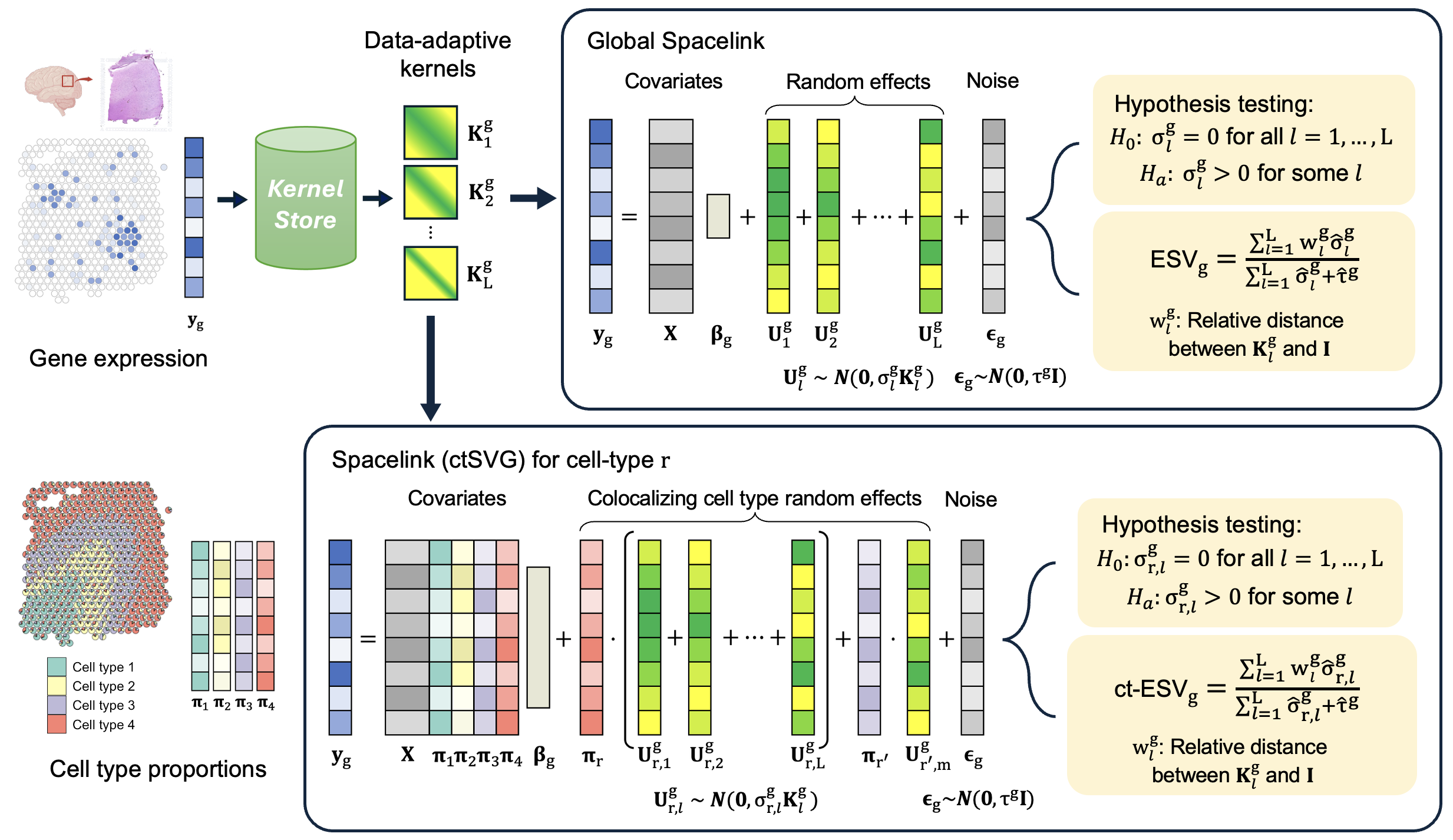
Spacelink is a unified statistical framework for detecting and prioritizing SVGs at both global tissue and cell-type resolution. Spacelink employs an adaptive multi-kernel model to capture spatial variance across diverse length scales, and its cell-type specific version introduces a data-driven gating strategy to correct for spatial colocalization, designed to improve the specificity for cell types that are weakly represented in mixed spots relative to more abundant colocalizing cell types. To summarize spatial variability, we define Effective Spatial Variability (ESV), a metric which integrates variance magnitude of each component kernel and its corresponding spatial scale into a single interpretable score directly suited for genetic analyses.
Installation
System requirements: - R ≥ 4.0.0 - Supported: Windows, macOS, Linux
You can install the development version of spacelink from GitHub with:
# install.packages("devtools")
devtools::install_github("hanbyul-lee/spacelink")The following dependency package may need to be installed manually from CRAN:
install.packages("gaston")The other dependency packages are as follows. If automatic installation fails, you can manually install them from CRAN:
install.packages(c("Rcpp", "RcppML", "pracma", "RANN", "fields", "Matrix"))Documentation
| Page | Description | Runtime |
|---|---|---|
| Installation | Setup | 3m |
| Quick Start | Get started with a minimal working example | 10s |
| Overview | Descriptions of main functions | - |
| Spacelink Global SVG Vignette | Examples and guides for Spacelink analysis at the global tissue level | 10s |
| Spacelink ct-SVG Vignette | Examples and guides for Spacelink (ct-SVG) analysis at the cell-type-specific level | 10s |
| Illustration on CosMx Data | Application of Spacelink on a large-scale single-cell resolution dataset | 50m |
| Illustration on Visium Data | Application of Spacelink on a medium-scale spot resolution dataset | 20m |
| Disease Informativeness Evaluation | Evaluation of ESV disease informativeness using PoPS (Polygenic Priority Score) | 2m |
| AD Data Analysis | Applying Spacelink to identify spatial gene expression changes in Alzheimer’s Disease | 1m |
| Lengthscale Estimation | Comparison of length-scale estimation performance | 5m |
| Runtime & Memory Usage | Benchmarks of computational time and memory requirements across different dataset sizes | 12m |